Making Sleep Mutants
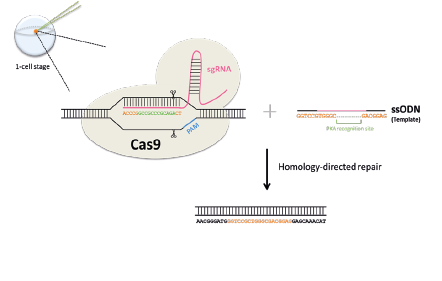
In zebrafish, it is possible to quickly target specific genes to make mutants using a system called CRISPR-Cas9. This is done by injecting into 1-cell stage zebrafish eggs a Cas9 protein along with a single guide RNA (sgRNA) that has a complementary sequence to the gene to target. The sgRNA will direct the Cas9 to cut the DNA at the gene of interest, creating a double-strand break.
When the cell repairs the break, it will occasionally make an error, leading to a mutation. If we supply a single stranded DNA oligonucleotide (ssODN) with homology to our targeted sequence, we can trick the cell's repair machinery into replacing the sequence with a specially edited version — for example one that is missing a few bases.
Mutations in Sleep Genes Lead to Changes in Behavior
After editing specific genes of interest, we track the zebrafish larvae over several days and nights to monitor their sleep and activity. The effects we observe in animals harbouring mutations can be quite specific to different times of the day.
For example, the trace on the right shows the effect of mutations in the zebrafish gene cntnap2, an autism risk gene. The cntnap2 mutant fish (red trace) are hyperactive compared to their unaffected siblings (blue), but only at night. By uncovering mutants with defects in different aspects of the timing and duration of behavior, we hope to unravel how sleep and wake are controlled. So far, we have found a variety of hyperactive and short sleeping mutants.
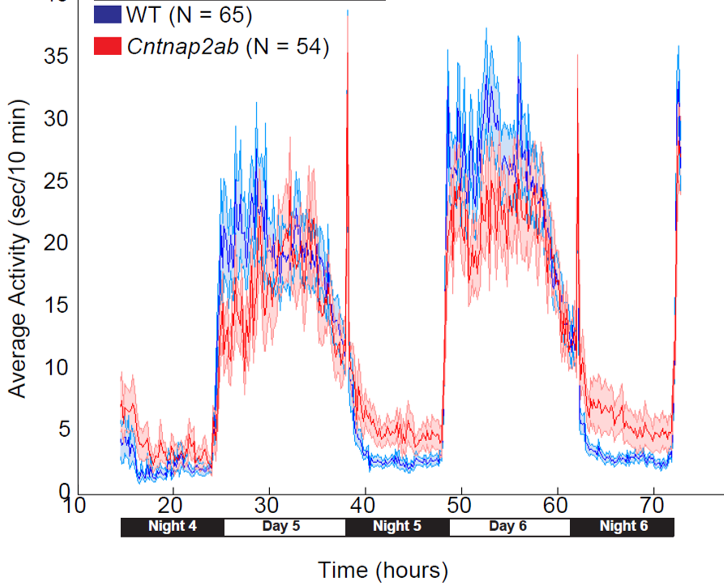
cntnap2 mutants are hyperactive at night. Red trace = mutant fish; blue trace = unaffected siblings.
Drug Screening
With our automated assay, we have generated a large dataset from the behavioral fingerprints of nearly 6,000 small molecules and identified hundreds of pharmacological agents that robustly altered zebrafish sleep — including not only known sedatives and psychotropic compounds but also novel molecules not previously implicated in sleep/wake regulation.
Some of the behavioral effects were unexpected. For example, a large class of ether-a-go-go related gene (ERG) potassium channel inhibitors increased wakefulness, but only at night. We also observed that a large panel of anti-inflammatory compounds increased wakefulness only during the day. More recently, we have demonstrated that the behavioral fingerprints representing drug effects on behavior can be used to predict small molecules that rescue mutant phenotypes — for example, the nighttime hyperactivity of the zebrafish autism model, cntnap2.
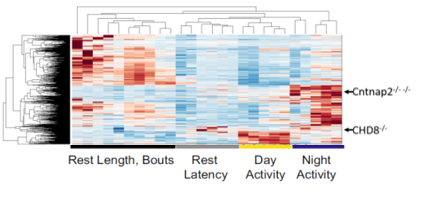
Mapping Mutants to a Drug Library
Each row in this figure represents a behavioral fingerprint for a drug that alters sleep in various ways. Red indicates a parameter that the drug increased, while blue indicates it was decreased. By mapping autism risk mutants (e.g. Cntnap2) to this dataset, we can identify drugs that rescue mutant sleep behaviors back to normal.