Linkage
mapping with phenotypic and molecular markers
Linkage
mapping is based on variation in crossover frequencies. Any locus for which
multiple alleles are available can be mapped. Visible phenotypic differences
between wild type and (usually recessive) mutant alleles can be easy to score
but are not available for all loci. DNA sequence polymorphisms (molecular
markers) are far more common. Many can, moreover, be identified relatively
efficiently by Southern blotting or PCR without sequencing the entire locus.
Consider
the simple dihybrid, or two point, cross that Morgan and Sturtevant devised to determine
the genetic distance between the Drosophila genes purple (pr) and vestigial (vg):
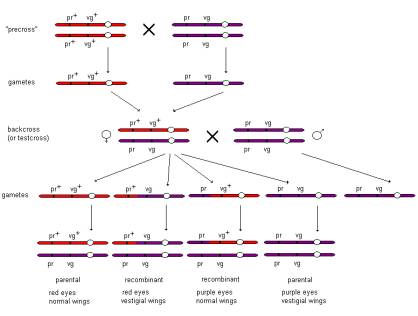
This
cross allows the relative distance between the two loci to be estimated from
the proportion of recombinant offspring ie offspring with a different combination
of eye colour (purple) and wing shape (vestigial) phenotypes than that seen in
either parent.
In this
case the simplest way to distinguish parental [pr/pr vg/vg] or [pr+/pr+ vg+/vg+] and recombinant [pr+/pr vg/vg] or [pr+/pr vg+/vg] flies is to look at them.
However, the wild type and mutant alleles of both genes also have differences
at the DNA sequence level and it would be theoretically possible to identify
parental and recombinant individuals by extracting DNA from each fly and
sequencing it. A severely colour blind geneticist could score each fly for wing
shape then extract the DNA from it to determine whether it were homozygous or
heterozygous for purple.
When mapping a gene in order to clone it, multi-point crosses are used to measure recombination frequencies between the locus being cloned and molecular markers for candidate genes or for known chromosomal locations. In classical linkage mapping tester stocks, homozygous mutant for several previously mapped loci are crossed to the new mutant. To facilitate mapping of the mutants generated by large scale screens the mutagenesis and subsequent breeding programmes are done in inbred lines, which are homozygous at a large number of RFLP and microsatellite loci. For mapping purposes pre-crosses and test crosses between fish carrying the mutation and a genetically distinct inbred strain are set up. This tester strain will have been selected to be homozygous for a different alleles of the RFLP and microsatellite markers carried by the mutant stock. Offspring generated by the test cross are scored for the mutant phenotype then DNA is extracted and can be analysed to determine whether any individual is homozygous or heterozygous for a suite of molecular markers.
References
http://www.ucl.ac.uk/~ucbhjow/b241/mapping_1.html
http://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=iga.section.899
http://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=iga.section.919