|
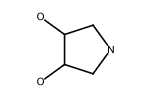
| Fragment 113 |
The PDB codes are hyperlinked to a schematic LIGPLOT diagram in PDBsum.
| Table S1.113. Fragment 113 interactions with Asp |
|
Listed below are the
ligands in the PDB that contain the chemical fragment shown on the
left and which interact via this fragment with the side chain of a
Asp residue via hydrogen bonds.
The PDB codes are hyperlinked to a schematic LIGPLOT diagram in PDBsum. |
| Ligand | Description | |||||||||||||||||||||||
|
|
1. Ligand: GB1 (2r,3r,4s)-2-({[(1r)-2-hydroxy-1-phenylethyl]amino}methyl)pyrrolidine-3,4-diol PDB code: 2f18. |
|||||||||||||||||||||||
|
|
2. Ligand: GB2 (2r,3r,4s)-2-({[(1s)-2-hydroxy-1-phenylethyl]amino}methyl)pyrrolidine-3,4-diol PDB code: 2f1a. |
|||||||||||||||||||||||
|
|
3. Ligand: GB3 (2r,3r,4s,5r)-2-({[(1r)-2-hydroxy-1-phenylethyl]amino}methyl)-5-methylpyrrolidine-3,4-diol PDB code: 2f1b. |
|||||||||||||||||||||||
|
|
4. Ligand: GHA 1'-((1,4-dideoxy-1,4-imino-d-arabinitol)-4-n-ammonium)-1'-deoxy-l-erythritol-3'-sulfate inner salt PDB code: 1tqu. |
|||||||||||||||||||||||
|
|
5. Ligand: IMU Phosphoric acid mono-[5-(2-amino-4-oxo-4,5-dihydro-3h-pyrrolo[3,2-d]pyrimidin-7-yl)-3,4-dihydroxy- pyrrolidin-2-ylmethyl] ester PDB codes: 1bzy, 1dqn. |
|||||||||||||||||||||||
|
|
6. Ligand: IRP (1s)-1(9-deazahypoxanthin-9yl)1,4-dideoxy-1,4-imino-d-ribitol-5-phosphate PDB code: 1cjb. |
|||||||||||||||||||||||
|
|
7. Ligand: W72 6-deoxy-6-[(2r,3r,4r)-3,4-dihydroxy-2-(hydroxymethyl)pyrrolidin-1-yl]-l-gulonic acid PDB code: 2fyv. |
|||||||||||||||||||||||
|
|
8. Ligand: IMH 1,4-dideoxy-4-aza-1-(s)-(9-deazahypoxanthin-9-yl)-d-ribitol PDB codes: 2ff1, 3b9g. |
|||||||||||||||||||||||
|
|
9. Ligand: IMQ (2r,3r,4s)-2-(hydroxymethyl)-1-(quinolin-8-ylmethyl)pyrrolidine-3,4-diol PDB code: 3epx. |
|||||||||||||||||||||||
|
|
10. Ligand: JMQ (2r,3r,4s)-2-(hydroxymethyl)-1-[(4-hydroxy-5h-pyrrolo[3,2-d]pyrimidin-7-yl)methyl]pyrrolidine-3,4- diol PDB code: 3epw. |
|||||||||||||||||||||||